Ancient pathogen meets new technology
London School of Hygiene & Tropical Medicine https://lshtm.ac.uk/themes/custom/lshtm/images/lshtm-logo-black.png Wednesday 3 February 2021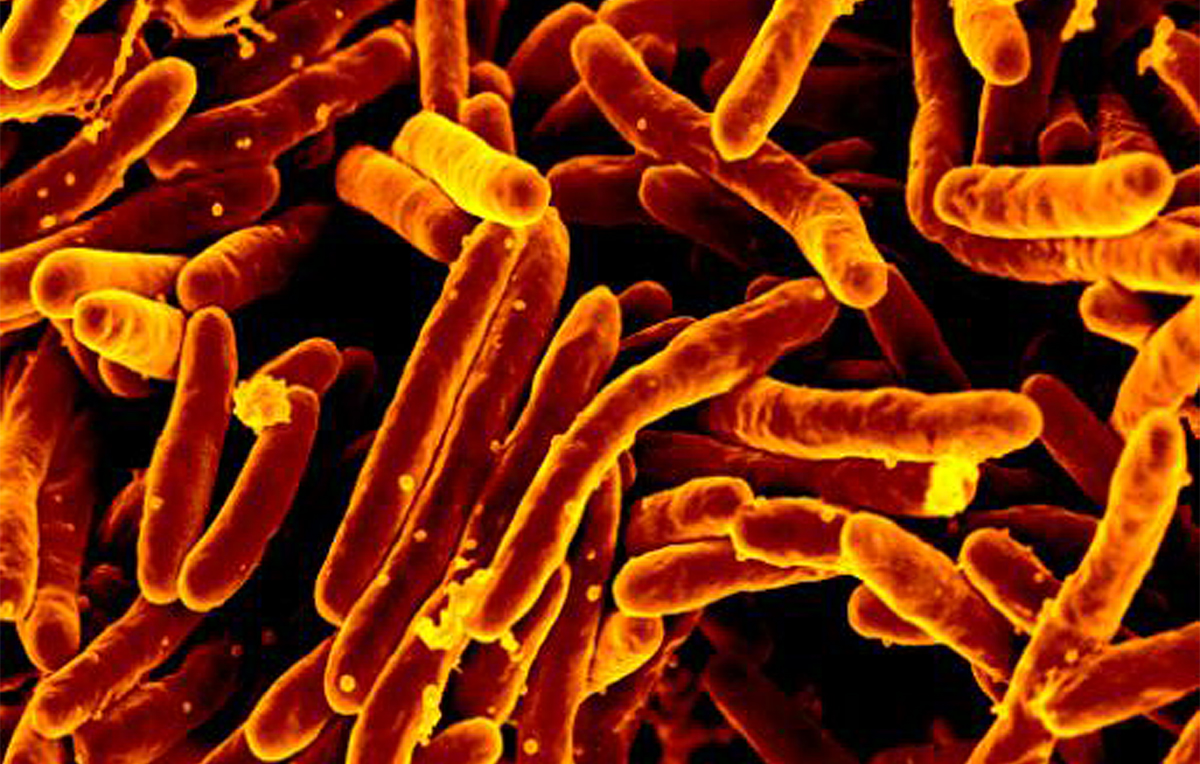
In the midst of the COVID-19 pandemic we are reminded that infectious diseases are not a thing of the past but a very real threat to millions of people globally. Tuberculosis (TB), caused by the bacterium Mycobacterium tuberculosis, is the biggest killer by a single bacterial pathogen, causing over 10 million people to fall ill and 1.5 million people to die from TB each year.
Although TB has been around for millennia (evidence of TB has been uncovered in ancient Egyptian mummies) there is still much we do not yet understand about either the disease or the organism that causes it. The reasons for this are complex: we lack animal models that accurately recapitulate the human disease and the bacteria that cause TB are slow-growing, highly pathogenic and have certain features that make them less amenable to traditional methods of study and manipulation. As a result, some of the specific adaptations and features that have made Mycobacterium tuberculosis such a successful human pathogen are yet to be fully explored and understood.
In our new study, published in Cell Reports, we applied cutting-edge biochemical techniques to examine one aspect of the fundamental biology of Mycobacterium tuberculosis. All organisms rely on protein molecules to control the cellular processes that keep them alive. Synthesising these proteins, which are encoded in the organism’s DNA (the genetic recipe book of the cell), is a highly regulated process with controls operating at every step.
We were interested in how particular signals or instructions within the genetic code contribute to the efficiency with which the protein is synthesised. During the research, we compared populations of bacteria that were growing normally with those that had been starved of nutrients for 24 hours, mimicking conditions the bacteria encounter in the human host.
We were able to reveal exactly which proteins were being made by the bacteria under each growth condition and how different signals in the DNA ‘recipe’ for the protein influence the process. This is important because it sheds light on some of the mechanisms by which bacteria are able to adapt to survival under stress conditions in the human host – and if we understand what they are doing to survive, we can intervene to prevent this and help clear the infection.
Not only did our study uncover important details about the fundamental life process of protein synthesis in Mycobacterium tuberculosis, which may lead to the development of novel therapeutics to target infection, it also demonstrated how new technologies can be applied to the study of an ancient pathogen. In contrast to research into COVID-19, development of better vaccines and treatments for TB has been slow, hindered in part by inherent difficulties in working with Mycobacterium tuberculosis.
Our study pushed the boundaries of techniques that could be adapted to study this pathogen – something that is desperately needed if we are to develop more targeted approaches to the prevention and cure of TB.
Publication
E. B. Sawyer, J. E. Phelan, T.G. Clark and T. Cortes. A snapshot of translation in Mycobacterium tuberculosis during exponential growth and nutrient starvation revealed by ribosome profiling. Cell Reports. DOI: 10.1016/j.celrep.2021.108695
LSHTM's short courses provide opportunities to study specialised topics across a broad range of public and global health fields. From AMR to vaccines, travel medicine to clinical trials, and modelling to malaria, refresh your skills and join one of our short courses today.