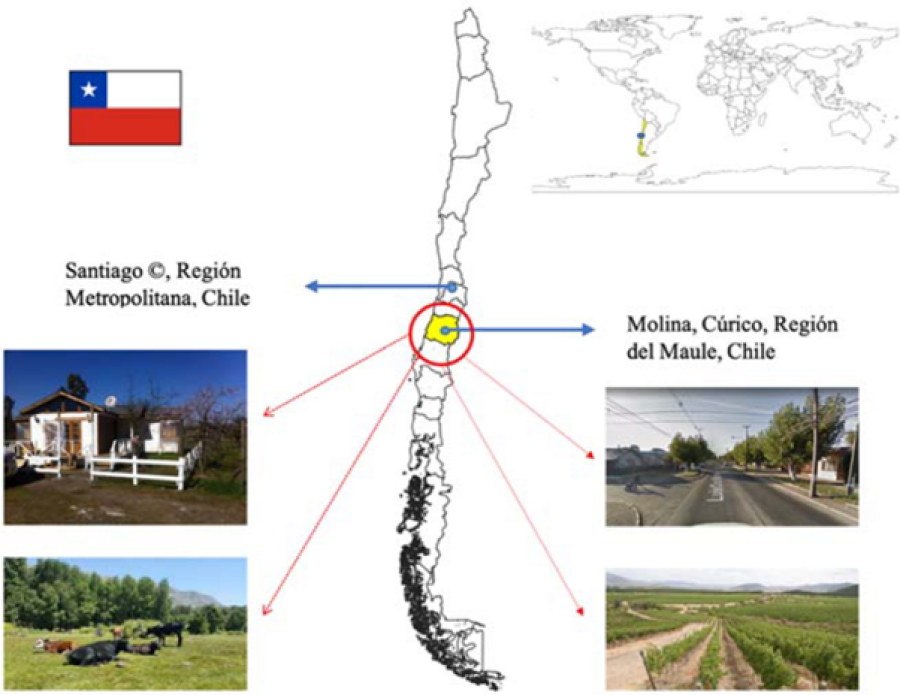
Biological specimen analysis and genomics studies are vital to understand AMR and its transmission patterns, but modelling too can provide extremely useful insights. Kasim Allel (PhD student at LSHTM) and colleagues developed a dynamic compartmental model to quantitatively characterise the dynamics of GN-ARB transmission in this community, among individuals at high and low risks of exposure to ARB reservoirs.
Their model suggests that 98% of the total GN-ARB acquisition/colonisation occurs in the community, compared to 1% in the hospital. Community acquired infections further accounted for 97.6% of blood stream infections in the model. Although transmission rates were higher in hospitals, the dissemination of pathogens, the number of colonised individuals and the disease burden was higher in the community due to the higher population size and contact with community ARB reservoirs. The main risk factors in the community were overcrowded spaces and households with higher animal contact, and previous antibiotic consumption.
How does this compare with biological analysis of community samples? Quite favourably as far as the environmental risk factors for human AMR are concerned.
A study from rural Ujjain in India, a low middle-income country, that looked at the associations between antibiotic resistance in humans, animals and different water sources in the same area over similar time points over two years, reported highest resistance prevalence in human samples, compared with animal and water samples. The mean of human cases attributable to each environmental source was highest for household drinking water (47.7), followed by animals (38.3) and waste water (36.7). It was the lowest for source drinking water (0.2), suggesting that resistance may have been related to how humans interacted with their drinking water in this setting, rather than the water source itself. Both studies highlighted the importance of the human-animal-environmental interface and the quintessential One Health approach for AMR, starting from community settings.
Active community surveillance programs can mitigate the potential risks of environmental and hospital exposures, contaminated food and zoonotic diseases among communities. However, in the absence of well-developed biological or modelling tools in low- and middle- income countries, how can epidemics resulting from AMR and One Health risk factors be predicted in future?
There is some evidence that futuristic epidemic intelligence could benefit from event-based surveillance besides indicator-based surveillance (e.g. from health facility or other reporting sites). Event based surveillance is performed through media monitoring using internet-based platforms with automated systems for scraping, filtering and analysing news as well as social media information. One WHO study reported that a quarter of infectious disease outbreaks in the WHO AFRO region between 2017-2020, were initially detected through event-based surveillance using media sources. An innovative health-tech start-up, SpatialApps, has gone one step ahead and combined events data with geospatial data (such as location of wet markets and climate risks) to identify early warning signs of zoonotic disease outbreaks in South Asia.
Further advances in innovative, reliable and cost-effective surveillance tools can identify and mitigate the risks of AMR and save lives worldwide.
If you enjoyed this article and would like to build a career in global health, we offer a range of MSc programmes covering health and data, infectious and tropical diseases, population health, and public health and policy.
Available on campus or online, including flexible study that works around your work and home life, be part of a global community at the UK's no.1 public health university.
